[Update 2022-10-18: Replace epsilon with delta in description of example
problem 7 in pgapack]
Many optimization problems involve constraints that valid solutions must
satisfy. A constrained optimization problem
is typically written as a nonlinear programming problem .
\begin{align*}
\hbox{Minimize} \; & f_i(\vec{x}), & i &= 1, \ldots, I \\
\hbox{Subject to} \; & g_j(\vec{x}) \le 0, & j &= 1, \ldots, J \\
& h_k(\vec{x}) = 0, & k &= 1, \ldots, K \\
& x_m^l \le x_m \le x_m^u, & m &= 1, \ldots, M \\
\end{align*}
In this problem, there are \(n\) variables (the vector
\(\vec{x}\) is of size \(n\)), \(J\) inequality constraints,
\(K\) equality constraints and the variable \(x_m\) must be in the
range \(|x_m^l, x_m^u|\) (often called a box constraint). The
functions \(f_i\) are called the objective functions. If there is more
than one objective function, the problem is said to be a multi
objective optimization problem as described previously in this blog .
In the following I will use some terms that were introduced in that
earlier blog-post without further explanation.
The objective functions are not necessarily minimized (as in the
formula) but can also be maximized if the best solutions to a problem
requires maximization. Note that the inequality constraints are often
depicted with a \(\ge\) sign but the formula can easily be changed
(e.g. by multiplying with -1) to use a \(\le\) sign.
Since it is very hard to fulfil equality constraints, especially if they
involve nonlinear functions of the input variables, equality constraints
are often converted to inequality constraints using an δ‑neighborhood:
\begin{equation*}
-\delta \le h_k(\vec{x}) \le \delta
\end{equation*}
Where δ is chosen according to the requirements of the problem
for which a solution is sought.
One very successful method of solving constrained optimization problems
is to consider a lexicographic ordering of constraints and objective
function values. Candidate solutions to the optimization problem are
first sorted by violated constraints (typically the sum over all
violated constraints) and then by the objective function value(s) .
When comparing two individuals during selection in the genetic algorithm
there are three cases: If both individuals violate constraints, the
individual with the lower constraint violation wins. If one violates
constraints and the other does not, the one not violating constraints
wins. In the last case where both individuals do not violate constraints
the normal comparison is used (which depends on the algorithm and if
we're minimizing or maximizing).
This method, originally proposed by Deb , is implemented in the
genetic algorithm package I'm maintaining, PGAPack, and my Python
wrapper PGAPy for it.
With this algorithm for handling constraints, the constraints are
optimized first before the algorithm "looks" at the objective
function(s) at all. It often happens that the algorithm ends up
searching in a region of the input space where no good solutions exist
(but no constraints are violated). Hard problems often contain equality
constraints (converted to inequality constraints as indicated earlier)
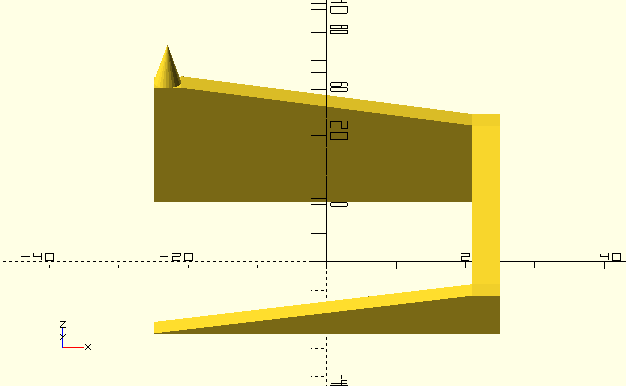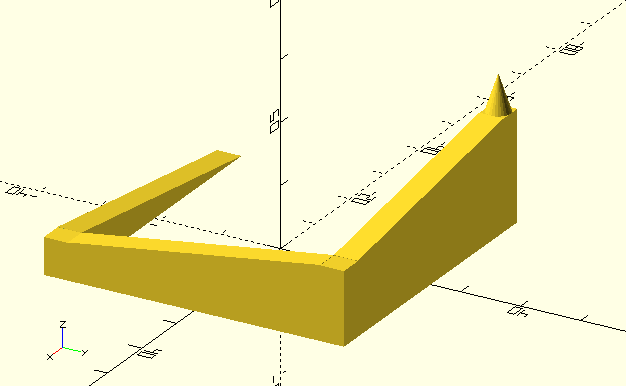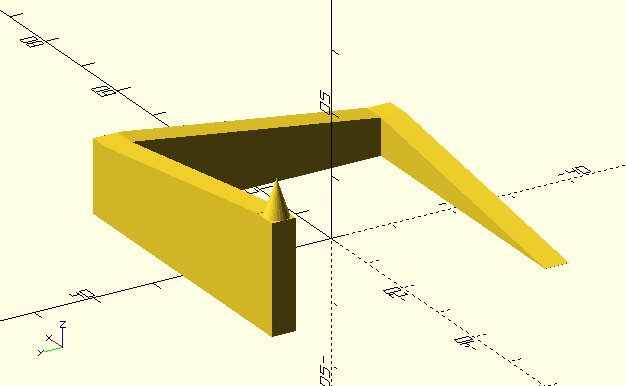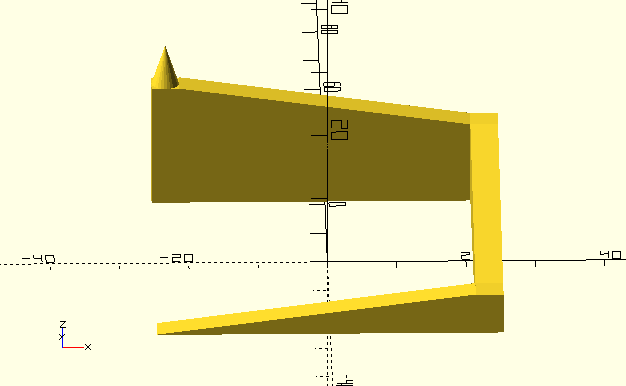
or other "hard" constraints. In my previous blog post on
antenna optimization I wrote: "the optimization algorithm has a hard
time finding the director solutions at all. Only in one of a handful
experiments I was able to obtain the pareto front plotted above".
In that experiment I was running 50 searches and only 5 of them did
not get stuck in a local optimum. A similar thing happens for the
problem (7) in Deb's paper which has equality constraints. I've
implemented this as example problem 7 in PGAPack. It only finds a
solution near the (known) optimum when \(\delta \ge 10^{-2}\) for
all equality constraints (I didn't experiment with different random
seeds for the optimizer, maybe a better solution would be possible with
a different random seed). In the paper , Deb uses
\(\delta = 10^{-3}\) for the same reason.
One method for handling this problem was appealing because it is so
easy to understand and implement:
Takahama and Sakai were first experimenting with a method for relaxing
constraints during the early stages of optization with a formulation
they called an α‑constrained genetic algorithm . They
later simplified the formulation and called the resulting algorithm
ε constrained optimization. It can be applied to different
optimization algorithms, not just genetic algorithms and variants .
Of special interest is the application of the method to differential
evolution , but of course it can also be applied to other forms
of genetic algorithms.
Note that the ε in the name of the algorithm can be used for
the δ used when converting an equality constraint to inequality
constraints but is not limited to this case.
During the run of the optimizer in each generation a new value for
ε is computed. The comparison of individuals outlined above is
modified, so that an individual is handled like it was not violating any
constraints if the constraint violation is below ε. So if both
individuals have constraint violations larger than ε, the one
with lower violation wins. If one violation is below ε and the
other above, the individual with the violation below ε wins. And
finally if the constraint violations of both individuals are below
ε, the normal comparison takes place.
The last case is the key to the success of this algorithm: Even though
the search proceeds into a direction where the constraint violations are
minimized, at the same time good solutions in terms of the objective
function are found.
The algorithm begins by initializing \(\varepsilon_0\) with the constraint
violation of the individuum with index \(\theta\) from the initial
population sorted by constraint violation, where
\(\theta\) is a parameter of the algorithm between 1 and the
population size, a good value uses the individuum at about 20%
of the population size which is also the default in PGAPack.
In each generation \(t\), \(\varepsilon_t\) is computed by
\begin{equation*}
\varepsilon_t = \varepsilon_0 \left(1-\frac{t}{T_c}\right)^{cp}
\end{equation*}
up to generation \(T_c\). After that generation, ε is set to
0. The exponent \(cp\) is between 2 and 10. The 2010 paper
recommends to set \(cp = 0.3 cp + 0.7 cp_\min\) at
generation \(T_\lambda = 0.95 T_c\) where \(cp_\min\) is
the fixed value 3. The initial value of \(cp\) is chosen so that
\(\varepsilon_\lambda=10^{-5}\) at generation \(T_\lambda\) unless
it is smaller than \(cp_\min\) in which case it is set to
\(cp_\min\). PGAPack implements this
schedule for \(cp\) by default but allows to change \(cp\)
at start and during run of the optimizer, so it's possible to easily
implement a different schedule for \(cp\) – the default works
quite nicely, though.
With the ε constraint method, example 7 from Deb can be
optimized with a precision of \(10^{-6}\) in my experiments, see the
epsilon_generation parameter in the optimizer example
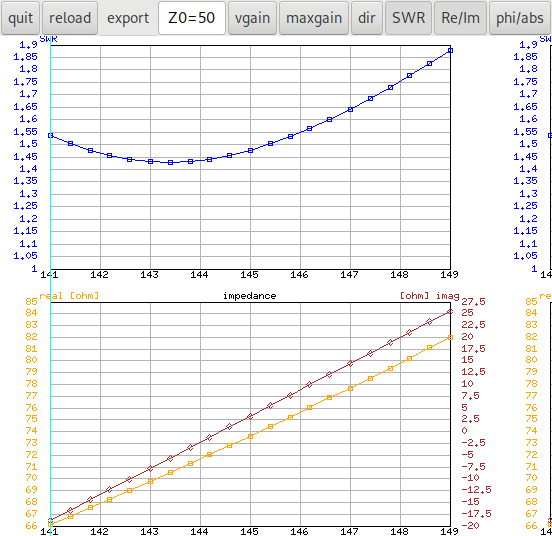
The antenna-optimizer with an ε‑generation of 50 (that's
the \(T_c\) parameter of the algorithm) gets stuck in the local
optimum only in one of 50 cases, all other cases find good results:
In that picture all the solutions that are dominated by solutions from
another run are drawn in black. It can be seen that the data from run
number 16 did not contribute any non-dominated solutions (on the right
side in the legend the number 16 is missing). You can turn off the
display of the dominated solutions by clicking on the black dot in the
legend.
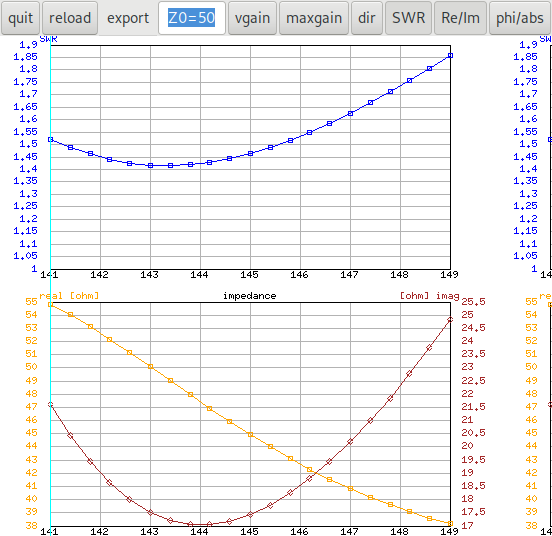
When I increase the ε‑generation to 60, the run with random
seed 16 also finds a solution:
We also see that the solutions are quite good (quite near to the pareto
front) for all runs, the black "shadow" of the dominated solutions is
quite near to the real pareto front and it is enough to do a single run
of the algorithm for finding a good set of solutions.